SPO-NDS – Swiss Personalized Oncology National data Stream
Prof. Olivier Michielin, MD-PhD and Sylvain Pradervand, PhD
Goal of the NDS:
The goal of the Swiss Personalized Oncology (SPO)-NDS is to build and extend the existing infrastructure, datasets, and know-how developed over the first 3 years of the Swiss Personalized Oncology SPHN Driver project. The SPO-NDS will aim to consolidate the clinical data acquisition, finalizing the extraction of the core data set (120 structured data entry types) at the 5 University Hospitals and at the Swiss Group for Clinical Cancer Research (SAKK), and pursuing the natural language processing (NLP) applications to enrich the data set. In addition, high value cohorts have been and will be assembled in cancer types treated with immuno-oncology (IO) therapies that will allow comparative immunomics between highly IO sensitive tumors (e.g., melanoma, lung-cancer, triple-negative breast cancer (TNBC)) to less sensitive (e.g., MSI-CRC, hormone receptor-positive breast cancer). Using the momentum of the Tumor Profiler (TuPro) initiative, single-cell and -omics data will now be at the center of the program including NGS, CyTOF/IMC, digital pathology, as well as functional drug screening to drive therapeutic decisions within the existing molecular tumor boards. The patients will readily benefit from these efforts as the infrastructure and the -omics data will be directly employed to support treatment decisions within the molecular tumor boards nationwide. In addition, tumors will be collected for subsequent analysis in a discovery program within the lighthouse project, using single-cell RNA seq, single-cell TCR seq, spatial transcriptomics, and bulk RNA seq. The lighthouse project will focus on identifying the mechanisms of primary and acquired immunotherapy resistance within and between tumors with different immune-reactivities. Our strategy is to use a comparative immune-oncology (IO) approach at the single-cell and multi-omics levels to identify shared features of primary and acquired resistance to IO in a subset of the molecular tumor board patients meeting strict inclusion criteria. In particular, we will compare the cancer-immune ecosystems of patients and tumors that respond well to IO to those who fail to mount antitumor immune responses to IO. In turn, these analyses are expected to identify predictive biomarkers to be used in precision oncology tumor boards, also directly benefiting patients.
Executive summary:
All aspects of the SPO-NDS will be geared towards their application in precision oncology molecular tumor boards at the local and national level, bringing unprecedented decision support for personalized therapies. As such, the SPO-NDS will create a direct link to patient care, allowing new treatment opportunities for patients who have escaped standard of care therapies or for whom several standard of care options exist without a rationale for selection. As the SPO core dataset will be used to track the patients enrolled in the molecular tumor boards, outcome measures for these treated patients will be channeled back into the dataset to further improve therapeutic decision support.
The SPO-NDS assembles medical oncologists specialized in precision oncology from the 5 university hospitals together with high profile scientists involved in specific -omics[1] analyses or in translational research in general. In order to maximize interactions between clinicians and scientists, the project is backed up by a solid IT infrastructure for clinical and -omics data exchange and interoperability as well as explicit interaction strategies within molecular tumor boards. The project aims to bring targeted -omics results back to patient care by ensuring national data interoperability, rapid turnaround times, and automated, dynamic, clinical reports so that the analyses can be used to fuel therapeutic decision making within the existing molecular tumor boards of the university hospital or the nationwide Swiss molecular tumor board initiated in the first part of the project supported by SPHN (SPO Driver project). As such, the SPO-NDS program will directly impact patient care. In addition, wider analyses will be conducted within the exploratory -omics stream on collected tumors. This second stream will not be directly connected to patient care as these more extended analyses are not currently compatible with clinical timelines. These data will be analyzed retrospectively using the most mature technology at the time of analysis, including scRNA seq, scTCR seq, and spatial transcriptomics. Such rich datasets will be used to fuel the lighthouse project where comparative immunomics will be central to the understanding of the factors associated with primary or acquired resistance in the selected cancer types (i.e., melanoma, lung-cancer, TNBC, MSI-CRC, or hormone receptor positive breast cancer), which display various levels of IO sensitivity. The analysis will aim at identifying predictive biomarkers to help treatment decisions, but also actionable pathways to design new treatment sequences or combinations.
Importantly, our comprehensive oncology infrastructure and rich datasets will be mutualized and reused in various Nested Projects like the study of the impact of germline genomics on IO sensitivity. As a second example of mutualization, we plan to make all IT infrastructures, SOPs, and analytics available to help develop a pediatric precision oncology program within our country.
Illustrative summary:
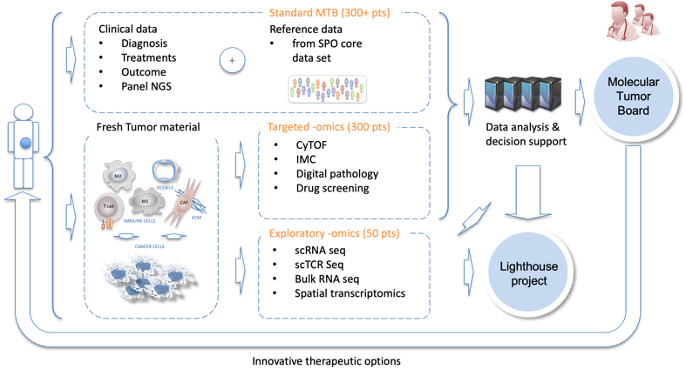
More about SPHN Grant:

Head of the Center for Precision Oncology
Head of Melanoma Clinic
Department of Oncology UNIL CHUV
Lausanne University Hospital and University of Lausanne, Switzerland.
Short Biography
Prof. Olivier Michielin obtained a Masters of Physics in 1991 at the Swiss Federal Institute of Technology and a Medical Degree from the University of Lausanne in 1997. He pursued his PhD training under the supervision of Jean-Charles Cerottini (Ludwig Institute) and Martin Karplus (Harvard and Strasbourg Universities, Chemistry Nobel Prize Laureate 2013). He was appointed Group Leader of the Swiss Institute of Bioinformatics in 2002 and became an Assistant Professor and Privat Docent at the Medical Faculty of Lausanne in 2004 and 2005, respectively. In parallel, he has trained as a medical oncologist and obtained his board certification in 2007 at the Department of oncology of the Lausanne University Hospital (CHUV) where he is currently heading the melanoma clinic. He was appointed Associate Professor in 2010 and full Professor in 2019. Prof. Olivier Michielin is mainly focused on translational oncology, developing new molecularly defined therapeutic approaches based on original bioinformatics techniques developed in his laboratory, as well as melanoma clinical trials at the Lausanne University Hospital. In 2016, Prof. Olivier Michielin was appointed Head of the Center for Precision Oncology within the Department of oncology.

Head of the Data Management and Analytics group at the Center for Precision Oncology led by Olivier Michielin
Short Biography
Sylvain Pradervand obtained a PhD degree in molecular biology at the University of Lausanne in 1998 under the supervision of Bernard Rossier. He then joined the laboratory of Prof. Kenneth R. Chien at the University of California San Diego (UCSD) to investigate the role of cytokines in heart diseases. With the emergence of genomic technologies, he turned his interests to bioinformatics and information technologies (professional certification in client-server technology from UCSD extension, certified Java programmer). He gained experience working for the bioinformatics team of the Alliance for Cellular Signaling (led by Nobel Prize Laureate Alfred Gilman) and participating in the development of the Signaling Gateway web site of the Nature Publishing Group. In 2006, he came back at the University of Lausanne as head of the bioinformatics unit of the Lausanne Genomic Technologies Facility (LGTF). In 2015, he joined the Lausanne University Hospital (CHUV) where he worked jointly with the Lausanne Institutional Biobank and the hospital IT department to set up the CHUV clinical research data warehouse. Since 2018, he is the head of the data management and analytics group at the Center for Precision Oncology led by Olivier Michielin.